Normalizing Flows¤
In this tutorial, adapted from this blog post, we will implement a simple RealNVP normalizing flow. Normalizing flows are a class of generative models which are advantageous due to their explicit representation of densities and likelihoods, but come at a cost of requiring computable jacobian determinants and invertible layers. For an introduction to normalizing flows, see Papamakarios et al. (2021).
References
RealNVP:
@inproceedings{dinh2017density,
title={Density estimation using {Real-NVP}},
author={Dinh, Laurent and Sohl-Dickstein, Jascha and Bengio, Samy},
booktitle={International Conference on Learning Representations},
year={2017},
}
Normalizing flows survey:
@article{papamakarios2021normalizing,
title={Normalizing Flows for Probabilistic Modeling and Inference},
author={Papamakarios, George and Nalisnick, Eric and Rezende, Danilo Jimenez
and Mohamed, Shakir and Lakshminarayanan, Balaji},
journal={Journal of Machine Learning Research},
volume={22},
number={57},
pages={1--64},
year={2021},
}
import equinox as eqx
import jax
import jax.numpy as jnp
import matplotlib.pyplot as plt
import optax
from sklearn import datasets
from sklearn.preprocessing import StandardScaler
from distreqx import bijectors, distributions
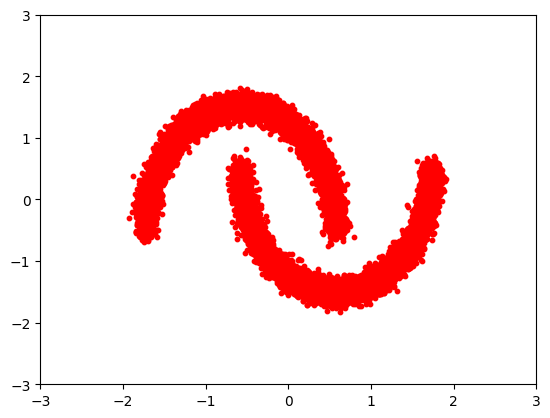
Let's define our simple dataset.
n_samples = 20000
noisy_moons = datasets.make_moons(n_samples=n_samples, noise=0.05)
X, y = noisy_moons
X = StandardScaler().fit_transform(X)
xlim, ylim = [-3, 3], [-3, 3]
plt.scatter(X[:, 0], X[:, 1], s=10, color="red")
plt.xlim(xlim)
plt.ylim(ylim)
(-3.0, 3.0)
Now we can program our custom bijector.
class RNVP(
bijectors.AbstractFwdLogDetJacBijector, bijectors.AbstractInvLogDetJacBijector
):
_is_constant_jacobian: bool
_is_constant_log_det: bool
sig_net: eqx.Module
mu_net: eqx.Module
d: int
k: int
flip: bool
def __init__(self, d, k, flip, key, hidden=32):
self._is_constant_jacobian = False
self._is_constant_log_det = False
self.flip = flip
keys = jax.random.split(key, 4)
self.d = d
self.k = k
self.sig_net = eqx.nn.Sequential(
[
eqx.nn.Linear(k, hidden, key=keys[0]),
eqx.nn.Lambda(jax.nn.swish),
eqx.nn.Linear(hidden, d - k, key=keys[1]),
]
)
self.mu_net = eqx.nn.Sequential(
[
eqx.nn.Linear(k, hidden, key=keys[2]),
eqx.nn.Lambda(jax.nn.swish),
eqx.nn.Linear(hidden, d - k, key=keys[3]),
]
)
def forward_and_log_det(self, x):
x1, x2 = x[: self.k], x[self.k :]
if self.flip:
x1, x2 = x2, x1
sig = self.sig_net(x1)
z1, z2 = x1, x2 * jnp.exp(sig) + self.mu_net(x1)
if self.flip:
z1, z2 = z2, z1
z_hat = jnp.concatenate([z1, z2])
log_det = jnp.sum(sig)
return z_hat, log_det
def inverse(self, y):
z1, z2 = y[: self.k], y[self.k :]
if self.flip:
z1, z2 = z2, z1
x1 = z1
x2 = (z2 - self.mu_net(z1)) * jnp.exp(-self.sig_net(z1))
if self.flip:
x1, x2 = x2, x1
x_hat = jnp.concatenate([x1, x2])
return x_hat
def forward(self, x):
y, _ = self.forward_and_log_det(x)
return y
def inverse_and_log_det(self, y):
raise NotImplementedError(
f"Bijector {self.name} does not implement `inverse_and_log_det`."
)
def same_as(self, other) -> bool:
return type(other) is RNVP
Since we want to stack these together, we can use a chain bijector to accomplish this.
n = 3
key = jax.random.key(0)
keys = jax.random.split(key, n)
bijector_chain = bijectors.Chain([RNVP(2, 1, i % 2, keys[i], 600) for i in range(n)])
Flows map p(x) -> p(z) via a function F (samples are generated via F^-1(z)). In general, p(z) is chosen to have some tractable form for sampling and calculating log probabilities. A common choice is Gaussian, which we go with here.
base_distribution = distributions.MultivariateNormalDiag(jnp.zeros(2))
base_distribution_sample = eqx.filter_vmap(base_distribution.sample)
base_distribution_log_prob = eqx.filter_vmap(base_distribution.log_prob)
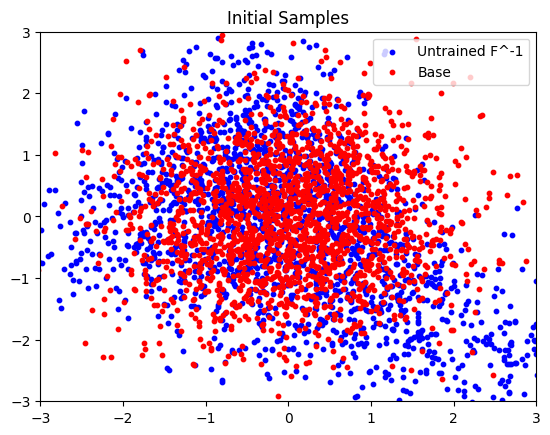
Here we plot the initial, untrained, samples.
num_samples = 2000
base_samples = base_distribution_sample(jax.random.split(key, num_samples))
transformed_samples = eqx.filter_vmap(bijector_chain.inverse)(base_samples)
plt.scatter(
transformed_samples[:, 0],
transformed_samples[:, 1],
s=10,
color="blue",
label="Untrained F^-1",
)
plt.scatter(base_samples[:, 0], base_samples[:, 1], s=10, color="red", label="Base")
plt.legend()
plt.xlim(xlim)
plt.ylim(ylim)
plt.title("Initial Samples")
plt.show()
learning_rate = 1e-3
num_iters = 1000
batch_size = 128
optimizer = optax.adam(learning_rate)
opt_state = optimizer.init(eqx.filter(bijector_chain, eqx.is_inexact_array))
def log_prob(params: bijectors.AbstractBijector, data):
f_inv, log_det = params.forward_and_log_det(data)
log_prob = base_distribution.log_prob(f_inv) + log_det
return log_prob
def loss(params, batch):
return -jnp.mean(eqx.filter_vmap(log_prob, in_axes=(None, 0))(params, batch))
@eqx.filter_jit
def update(model, batch, opt_state, optimizer):
val, grads = eqx.filter_value_and_grad(loss)(model, batch)
update, opt_state = optimizer.update(grads, opt_state)
model = eqx.apply_updates(model, update)
return model, opt_state, val
losses = []
for i in range(num_iters):
if i % 500 == 0:
print(i)
batch_indices = jax.random.choice(
key, jnp.arange(X.shape[0]), (batch_size,), replace=False
)
batch = X[batch_indices]
bijector_chain, opt_state, loss_val = update(
bijector_chain, batch, opt_state, optimizer
)
losses.append(loss_val)
0
500
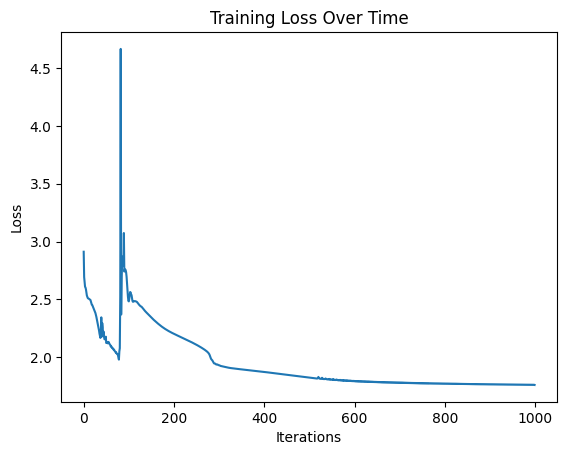
plt.plot(losses)
plt.xlabel("Iterations")
plt.ylabel("Loss")
plt.title("Training Loss Over Time")
plt.show()
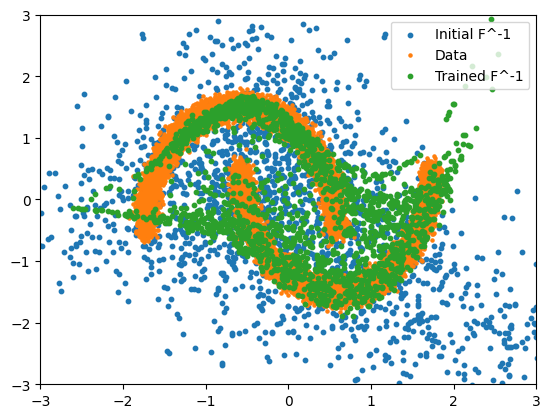
After training we can plot both F(x) (to see where the true data ends up in our sampled space) and F^-1(z) to generate new samples.
trained_params = bijector_chain
transformed_samples_trained = eqx.filter_vmap(bijector_chain.inverse)(base_samples)
plt.scatter(
transformed_samples[:, 0], transformed_samples[:, 1], s=10, label="Initial F^-1"
)
plt.scatter(X[:, 0], X[:, 1], s=5, label="Data")
plt.scatter(
transformed_samples_trained[:, 0],
transformed_samples_trained[:, 1],
s=10,
label="Trained F^-1",
)
plt.xlim(xlim)
plt.ylim(ylim)
plt.legend()
plt.show()
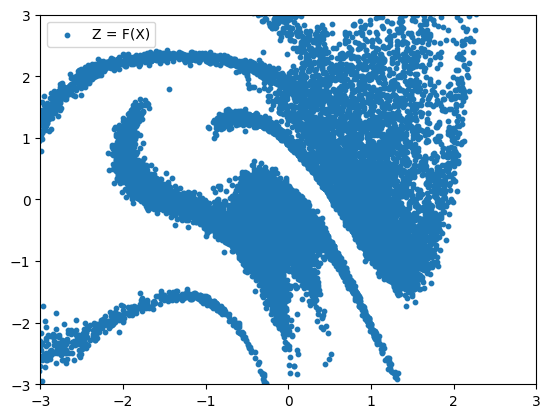
data_to_noise = eqx.filter_vmap(bijector_chain.forward)(X)
plt.scatter(data_to_noise[:, 0], data_to_noise[:, 1], s=10, label="Z = F(X)")
plt.xlim(xlim)
plt.ylim(ylim)
plt.legend()
plt.show()